Search SDAP
SDAP All
SDAP Food
SDAP Tools
AllergenAI (New!)
FAO/WHO Allergenicity Test
FASTA Search in SDAP
Peptide Match
Peptide Similarity
Peptide-Protein PD Index
Aller_ML, Allergen Markup Language
List SDAP
About SDAP
General Information
Manual
FAQ
Publications
Who Are We
Advisory Board
New Allergen Submission form
Our Software Tools
MPACK
FANTOM
GETAREA
InterProSurf
EpiSearch
Allergen Databases
WHO/IUIS Allergen Nomenclature database
FARRP Allergen Protein Database (University of Nebraska)
Allergen Database for Food Safety (ADFS)
COMPARE database
ALLFAM (Medical University of Vienna)
Allermatch (Wageninen University)
Allergome Database
Protein Databases
PDB
MMDB - Entrez
SWISS-PROT
NCBI - Entrez
PIR
Protein Classification
CATH
FSSP
iProClass
ProtoMap
SCOP
VAST
Bioinformatics Servers
BLAST @ NCBI
FASTA @ EMBL-EBI
Peptide Match @ PIR
ClustalW @ EMBL - EBI
SDAP 2.0 - Structural Database of Allergenic Proteins
| Go to: SDAP All allergens | Go to: SDAP Food allergens | |
| Send a comment to Werner Braun | Submit new allergen information to SDAP | |
| Alphabetical listing of allergens: A B C D E F G H I J K L M N O P Q R S T U V W X Y Z | ||
Access to SDAP is available
free of charge for Academic and non-profit use.
Licenses for commercial use can be obtained by contacting W. Braun (webraun@utmb.edu).Secure access to SDAP is available from https://fermi.utmb.edu/SDAP | ||
|
SDAP is a Web server that integrates a database of allergenic proteins with various computational tools that can assist structural biology studies related to allergens. SDAP is an important tool in the investigation of the cross-reactivity between known allergens, in testing the FAO/WHO allergenicity rules for new proteins, and in predicting the IgE-binding potential of genetically modified food proteins. Using this Internet service through a browser, it is possible to retrieve information related to an allergen from the most common protein sequence and structure databases (SwissProt, PIR, NCBI, PDB), to find sequence and structural neighbors for an allergen, and to search for the presence of an epitope other the whole collection of allergens. Read an SDAP Overview or select a SDAP function from the left column. |
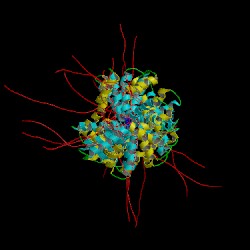
Structure of Ole e 6 - PDB 1SS3 |
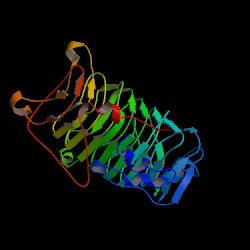
Structure of Jun a 1 - PDB 1PXZ |
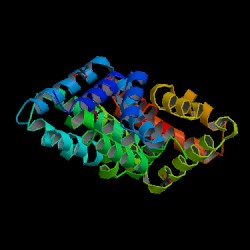
Structure of Fel d 1 - PDB 1PUO |
If SDAP is used in publications, please cite:
- Negi, S.S., Schein C.H., and Braun, W., The updated Structural Database of Allergenic Proteins (SDAP 2.0) provides 3D models for allergens and incorporated Bioinformatics Tools. Journal of Allergy and Clinical Immunology: Global (Volume 2, Issue 4, November 2023, 100162) [Abstract] [Full Paper]
-
Schein C.H.,Negi, S.S., and Braun, W.,
Still SDAPing Along: 20 Years of the Structural Database of Allergenic Proteins
Front. Allergy 3:863172. doi: 10.3389/falgy.2022.863172
[Pubmed]
[Full Paper]
- Ivanciuc, O., Schein. C. H., and Braun, W. SDAP: Database and Computational Tools for Allergenic Proteins. Nucleic Acids Res. 2003. 31(1). 359-362. [PubMed] [Full Paper]
- Ivanciuc, O., Schein. C. H., and Braun, W. Data Mining of Sequences and 3D Structures of Allergenic Proteins. Bioinformatics 2002, 18(10), 1358-1364. [PubMed] [Full Paper]
More recent publications related to SDAP, please cite:
- Negi, S.S., Goldblum, R., Braun, W., and Horiuti, T.M., Design of peptides with high affinity binding to a monoclonal antibody as a basis for immunotherapy Peptides. 145:170628, 2021 [PubMed] [Full Paper]
- Dreskin, S.C., Koppelman, S.J., Andorf, S., Nadeau, K.C., Kalra, A., Braun, W., Negi, S.S., Chen, X., Schein C.H., The importance of the 2S albumins for allergenicity and cross-reactivity of peanuts, tree nuts, and sesame seeds Journal of Allergy and Clinical Immunology 147 (4), 1154-1163, 2021. [PubMed] [Full Paper]
- Dreskin S.C , Germinaro M., Reinhold D., Chen X., Vickery B.P., Kulis M., Burks W.A., Negi S.S., Braun W., Chambliss J.M., Eglite S., and McNulty C.M.G, IgE binding to linear epitopes of Ara h 2 in peanut allergic preschool children undergoing oral Immunotherapy Pediatric Allergy and Immunology 30 (8), 817-823, 2020 [PubMed] [Full Paper]
- Lu, W., Negi, S.S., Schein, C.H., Maleki, S.J., Hulburt, B.K. and Braun, W., Distinguishing allergens from non-allergenic homologues using Physical-Chemical Property (PCP) motifs, Molecular Immunology, 9, 1-8, 2018. [PubMed] [Full Paper]
- Negi, S.S., and Braun, W., Cross-React: a new structural bioinformatics method for predicting allergen cross-reactivity, Bioinformatics, 33(7):1014-1020, 2017. [PubMed] [Full Paper]
- Power, T.D, Ivanciuc, O., Schein, C.H. and Braun, W. Assessment of 3D models for allergen research. PROTEINS: Struct. Funct. and Bioinf., 81(4), 545-554, 2013 [PubMed] [Full Paper]
- Ivanciuc, O., Gendel, S.M., Power, T.D., Schein, C.H. and Braun, W. AllerML: Markup language for allergens. Regul. Toxicol. Pharmacol., 60(1), 151-160, 2011. [PubMed] [Full Paper]
- Ivanciuc, O., Schein, C.H., Garcia, T.I., Oezguen, N., Negi, S.S. and Braun, W. Structural analysis of linear and conformational epitopes of allergens. Regul. Toxicol. Pharmacol., 54(3 Suppl), S11-S19, 2009. [PubMed] [Full Paper]
- Schein, C.H., Ivanciuc, O., Midoro-Horiuti, T., Goldblum, R.M. and Braun, W. An allergen portrait gallery: representative structures and an overview of IgE binding surfaces. Bioinform. Biol. Insights, 4, 113-125, 2010. [PubMed] [Full Paper]
We use the IUIS nomenclature and allergens in SDAP as the official set of allergens, as these allergens have been reviewed by a committee of experts in the field. However, we also included other proteins in SDAP when they were listed in an allergen data base at the time of establishing SDAP. We keep these non-IUIS allergens in SDAP (clearly marked as non-IUIS) as a service for the allergen researchers who are exploring and studying proteins that might have a potential allergenic response. In most cases these proteins can be searched in literature data bases for additional information.
The SDAP project is supported by grants from the National Institute of Health (2R56AI064913), the U.S. Environmental Protection Agency under a STAR Research Assistance Agreement (No. RD 834823), the National Institute of Health (1RO1AI165866-01), and by the Margaret Maccallum Gage and Tracy Davis Gage foundation.
Access to SDAP is available free of charge for Academic and non-profit use. Licenses for commercial use can be obtained by contacting W. Braun (webraun@utmb.edu).
Recent SDAP developments:
- PD distance: Click here to download PD Distance Perl Script , PD Distance python Script written by Surendra Negi.
- PeptideCutter@ExPASy: Protease cleavage sites predicted with PeptideCutter from ExPASy
- Allergen classification with Superfamily
- Allergen classification with InterPro
- List of 1908 allergens and isoallergens: List
- List of 1628 Protein sequences for allergens and isoallergens: List
- 109 Allergens with PDB structures: List
- 378 3D models for allergens and isoallergens: List
- 30 Allergens with IgE epitope sets: List
- 233 new Pfam allergen classes: List
- Implementation of the FAO/WHO Allergenicity Test
- FASTA search against all SDAP allergens
- Compute the sequence similarity for two sequences provided by the user using the PD index
- Search with a user-provided epitope (peptide) for similar regions in all SDAP allergens , using our original PD sequence similarity index
Previous version of SDAP was developed by Dr. Ovidiu Ivanciuc at The University of Texas Medical Branch, Galvestion, TX. Current version of SDAP is developed by Dr. Surendra Negi at the The University of Texas Medical Branch, Galvestion,TX.
