
SDAP Home Page
SDAP Overview
SDAP All
SDAP Food
Use SDAP All
Search SDAP All
List SDAP All
SDAP Food
SDAP Tools
FAO/WHO Allergenicity Test
FASTA Search in SDAP
Peptide Match
Peptide Similarity
Peptide-Protein PD Index
Aller_ML, Allergen Markup Language
List SDAP
About SDAP
General Information
Manual
FAQ
Publications
Who Are We
Advisory Board
Allergy Links
Our Software Tools
MPACK
FANTOM
GETAREA
NOAH/DIAMOD
MASIA
PCPMer
InterProSurf
EpiSearch
Protein Databases
PDB
MMDB - Entrez
SWISS-PROT
NCBI - Entrez
PIR
Protein Classification
CATH
CE
FSSP
iProClass
ProtoMap
SCOP
TOPS
VAST
Bioinformatics Servers
@TOME
BLAST @ NCBI
BLAST @ PIR
FASTA @ PIR
Peptide Match @ PIR
ClustalW @ BCM
ClustalW @ EMBL - EBI
ClustalW @ PIR
Bioinformatics Tools
Cn3D
MolMol
Bioinformatics Links
Bioinformatics Links Directory
|
SDAP - All Allergens
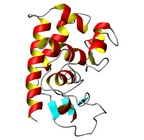 | Allergen Asp f 3Translate to AllerML | 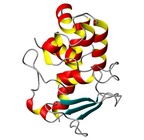 | 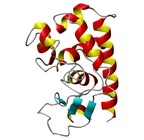 | | Allergen | Asp f 3 | | Type | fungi (moulds) | | Species - Systematic Name | Ascomycota Eurotiales; Aspergillus fumigatus | | Species - Common Name | | | Keywords | peroxisomal protein; Thioredoxin reductase; PMP20; Putative peroxiredoxin; EC 1. | | Class | IUIS |
| 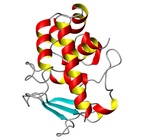 |
| Asp f 3 - PubMed | Reference | | Reference 1 | Ramachandran H, Jayaraman V, Banerjee B, Greenberger PA, Kelly KJ, Fink JN, Kurup VP, IgE binding conformational epitopes of Asp f 3, a major allergen of Aspergillus fumigatus. Clin Immunol. 2002 Jun;103(3 Pt 1):324-33. | | Reference 2 | Hemmann S, Ismail C, Blaser K, Menz G, Crameri R, Skin-test reactivity and isotype-specific immune responses to recombinant Asp f 3, a major allergen of Aspergillus fumigatus. Clin Exp Allergy. 1998 Jul;28(7):860-7. |
Asp f 3 - Protein Sequences
| Source | Link to Source | View Sequence | FASTA@SDAP< |
|

