
SDAP Home Page
SDAP Overview
SDAP All
SDAP Food
Use SDAP All
Search SDAP All
List SDAP All
SDAP Food
SDAP Tools
FAO/WHO Allergenicity Test
FASTA Search in SDAP
Peptide Match
Peptide Similarity
Peptide-Protein PD Index
Aller_ML, Allergen Markup Language
List SDAP
About SDAP
General Information
Manual
FAQ
Publications
Who Are We
Advisory Board
Allergy Links
Our Software Tools
MPACK
FANTOM
GETAREA
NOAH/DIAMOD
MASIA
PCPMer
InterProSurf
EpiSearch
Protein Databases
PDB
MMDB - Entrez
SWISS-PROT
NCBI - Entrez
PIR
Protein Classification
CATH
CE
FSSP
iProClass
ProtoMap
SCOP
TOPS
VAST
Bioinformatics Servers
@TOME
BLAST @ NCBI
BLAST @ PIR
FASTA @ PIR
Peptide Match @ PIR
ClustalW @ BCM
ClustalW @ EMBL - EBI
ClustalW @ PIR
Bioinformatics Tools
Cn3D
MolMol
Bioinformatics Links
Bioinformatics Links Directory
|
SDAP - All Allergens
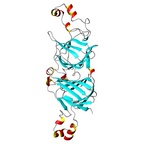 | Allergen Ara h 1Translate to AllerML | 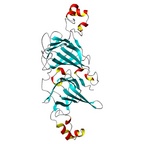 | 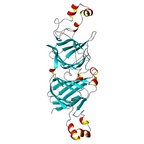 | | Allergen | Ara h 1 | | Type | foods | | Species - Systematic Name | Arachis hypogaea | | Species - Common Name | peanut | | Keywords | Vicilin; Clone P17
| | Class | IUIS |
| 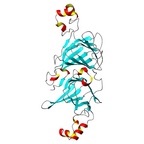 |
| Ara h 1 - PubMed | Reference | | Reference 1 | Burks AW, Cockrell G, Stanley JS, Helm RM, Bannon GA. Recombinant peanut allergen Ara h I expression and IgE binding in patients with peanut hypersensitivity.J Clin Invest. 1995 Oct;96(4):1715-21. | | Reference 2 | Kolarich D, Altmann F. N-Glycan analysis by matrix-assisted laser desorption/ionization mass spectrometry of electrophoretically separated nonmammalian proteins: application to peanut allergen Ara h 1 and olive pollen allergen Ole e 1. Anal Biochem. 2000 Oct 1;285(1):64-75. |
Ara h 1 - Protein Sequences
| Source | Link to Source | View Sequence | FASTA@SDAP | BLAST@NCBI | |
|

